- Review
- Published:
The AAA+ superfamily of functionally diverse proteins
Genome Biology volume 9, Article number: 216 (2008)
Summary
The AAA+ superfamily is a large and functionally diverse superfamily of NTPases that are characterized by a conserved nucleotide-binding and catalytic module, the AAA+ module. Members are involved in an astonishing range of different cellular processes, attaining this functional diversity through additions of structural motifs and modifications to the core AAA+ module.
The 'ATPases associated with diverse cellular activities' (AAA+ proteins) form a large and diverse superfamily found in all organisms. These proteins typically assemble into hexameric ring complexes that are involved in the energy-dependent remodeling of macromolecules [1]. Members of the AAA+ superfamily contain a highly conserved ATPase module of 200-250 amino acids, which includes an αβα core domain where the Walker A and B motifs of the P-loop NTPases are found [2–4].
AAA+ proteins are involved in a wide variety of different functions in which the energy extracted from ATP hydrolysis is used in molecular remodeling events. They are involved in processes as diverse as protein unfolding and degradation, peroxisome biogenesis, bacteriochlorophyll biosynthesis, and DNA recombination, replication and repair. AAA+ proteins include the molecular motor dynein, helicases involved in DNA replication, metal chelatases, and proteasome-associated proteins. As a consequence of their diverse functions, AAA+ proteins can be found in most subcellular compartments of eukaryotic cells, as well as in archaea, bacteria and viruses (Table 1). Interestingly, there is little correlation found between the clade an AAA+ protein belongs to and a specific remodeling activity. This suggests that the evolution of AAA+ proteins involved the initial emergence of a small number of defined AAA+ clades that, subsequently, expanded and adapted to allow the processing of a wide variety of targets. Furthermore, the emergence of partner proteins and cofactors has increased the functional diversity of AAA+ proteins [1, 5].
Structure and classification
Classification
Sequence and structure analyses reveal that the AAA+ superfamily underwent considerable divergence both before and since the appearance of the last common ancestor of the bacterial, archaeal and eukaryotic divisions of life [1, 3, 6]. Phylogenetic studies based on sequence and structural information divide the AAA+ superfamily into defined groups, clades and families [3, 5, 6]. The clades within each group are differentiated on the basis of the presence of distinct structural elements within and around the core AAA+ fold. This classification highlights the fact that many of these AAA+ lineages have evolved along different routes to acquire their unique functional differences.
The different clades fall within five major groups as shown in Table 1. These are: the extended AAA group; the helicases and clamp loaders (HEC) group; the protease, chelatase, transcriptional activators, and transport (PACTT) group, the ExeA group, and the signal transduction ATPases with numerous domains (STAND) group. Members of each of the major groups within the AAA+ superfamily can be found in all three of the major domains of life, with the exception of the ExeA group, which has so far only been detected in bacteria (Table 1).
Characteristic structural features
The AAA+ superfamily falls within the second major structural group of the P-loop NTPases, referred to as additional strand catalytic E (ASCE) [7]. As with all P-loop NTPases, members of this group possess a core αβα nucleotide-binding domain which contains two major nucleotide-binding and hydrolysis motifs referred to as Walker A (the P-loop) and Walker B. ASCE members are, however, distinguished from the other major P-loop structural group (kinase-GTPase or KG) by a characteristic 51432 order of β-strands in the β-sheet and the presence of a catalytic glutamate (E) residue within the Walker B motif [8] (Figure 1a,b).
Structure of the AAA+ module. (a) Monomeric AAA+ module of Aquifex aeolicus DnaA, a protein involved in the initiation of DNA replication (Protein Data Bank (PDB) code 2HCB) [5]. The α-helices and random coils are in green and the β-strands of the core αβα nucleotide-binding domain are in blue, with the exception of the two equal-sized helical inserts, which are colored pink. The small α-helical domain is colored purple. (b) Major motifs in the AAA+ module of (a) are colored as indicated in the key, on the basis of the alignment in reference [3]. The bound adenosine 5'-[β,γ-methylene]triphosphate (β,γ-methylene-ATP, a nonhydrolyzable ATP analog, orange sticks) and Mg2+ (black sphere) are also shown. (c) Top and side views of the hexameric structure of Haemophilus influenzae HslU, a member of the HslU/ClpX family (PDB 1KYI) [64]. α-Helices, including random coils, and β-strands of the core αβα nucleotide-binding domain are colored green and blue, respectively. Two additional helices characteristic of HslU-family proteins, called the I domain, are colored orange, and an additional extended loop between the second core β-strand and the following helix is colored in pink. The core small α-helical domain is colored purple, with the two-stranded β-sheet insertion in yellow. Structures were drawn using PyMOL [65].
AAA+ proteins, like many other members of the ASCE structural group, typically function as oligomeric rings, with a hexameric arrangement being most common (Figure 1c) [1]. In addition to the core features of the ASCE group, members of the AAA+ superfamily also contain a number of other sequence and structural characteristics, which serve to define their lineage within the ASCE group as a whole, as well as within the AAA+ superfamily itself.
The defining features of the AAA+ proteins can all be found within a region of 200-250 amino acids, generally referred to as the 'AAA+ module' [3, 4]. This module is comprised of two distinct domains: a core αβα nucleotide-binding domain and a smaller α-helical domain consisting of two helical hairpins arranged in a left-handed, superhelical structure (Figure 1a). The latter domain is poorly conserved at the sequence level, but is highly conserved structurally and serves as a defining characteristic of all AAA+ superfamily members [4, 6].
Within the AAA+ module there are numerous distinct signature sequences (Figure 1b). The Walker A motif (consensus Gx2GxGK [S/T], where G is glycine, K is lysine, S is serine, T is threonine, and x is any residue), lies between the first strand of the core β-sheet and the following helix, and plays an important role in nucleotide binding and metal-ion coordination [2, 5, 9]. The Walker B motif, (consensus ϕ4DE, where ϕ is a hydrophobic residue, D is aspartate and E is glutamate), is associated with the third strand of the core β-sheet, and contains residues involved in ATP hydrolysis and metal-ion coordination [2, 8].
The AAA+ module also contains a number of motifs that are not characteristic of P-loop NTPases as a whole, including sensor 1, sensor 2, and 'box' sequences (Figure 1b) [1, 3]. Sensor 1 is found on the fourth core β-strand and is characterized by a conserved polar residue, generally asparagine, threonine or histidine. The motif is critically important for the proteins' function and is proposed to interact either directly with the γ-phosphate of ATP, acting as a sensor of nucleotide binding/hydrolysis, or indirectly, via a water molecule, possibly helping to properly orient the water for nucleophilic attack on the bound nucleotide substrate [10, 11]. Sensor 2 maps to the third helix of the small α-helical domain and contains a conserved arginine residue. This residue interacts with the γ-phosphate of bound ATP substrate and is associated with a range of different roles, including nucleotide binding/hydrolysis and both inter- and intrasubunit communication and movement [3, 4, 12]. 'Box' motifs include Box II, which maps to the first helix before the core β-sheet and may be involved in adenine recognition; Box VII, which is located at the amino terminus of the fifth β-strand and contains an arginine finger that interacts with nucleotide bound by a neighboring subunit and is believed to play a role in ATP hydrolysis and intersubunit communication; and Boxes IV, IV', VII' and VII" [3, 13].
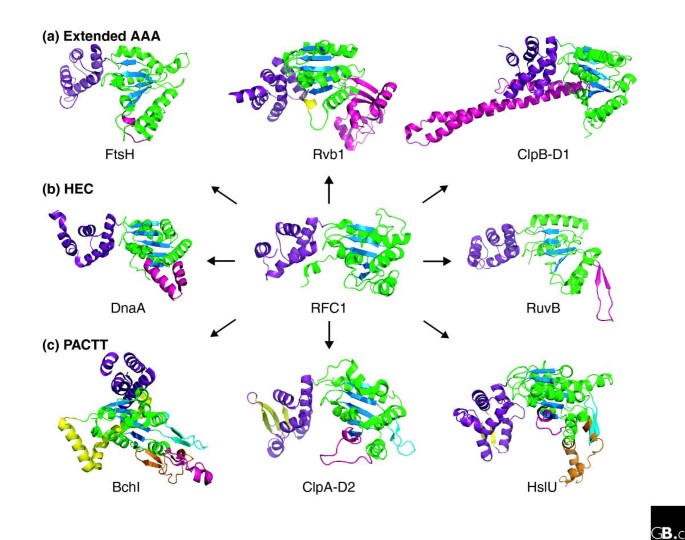
Different clades and families within the AAA+ superfamily also display unique structural modifications to the core AAA+ module, many of which possibly play a role in directing these families towards specific functions. Members of the clamp loader clade of the HEC group (see Table 1) best represent the 'basic' or 'core' AAA+ module, generally containing little to no modification [1]. The clamp loaders serve as mobile structures to which DNA polymerase core enzyme can be mounted during the process of DNA replication. The RFC1 protein of Saccharomyces cerevisiae shown in Figure 2 exemplifies this clade. Figure 2 also shows the AAA+ module structures of selected members of different clades/families, highlighting distinct structural modifications.
Structures of the AAA+ modules of selected superfamilymembers (see Table 1). The core αβα nucleotide-binding domains are shown in green (α-helices and random coil) and blue (β-strands). The small, α-helical domain of each AAA+ module is shown in purple. The canonical AAA+ module structure is exemplified by that of RFC1, which is shown in the center. (a) Representative members of the extended AAA group [6]. The FtsH AAA+ module from Thermus thermophilus (left, PDB 2DHR) contains an additional small helix (pink) downstream of the second β-strand, which is characteristic of the classical AAA clade [1,15]. The function of FtsH is discussed in the text. The Rvb AAA+ module, represented by human Rvb1 (center, PDB 2C90), contains a β-sheet-rich insert (pink) upstream of the Walker B motif and an additional small helix (yellow) downstream of the second β-strand of the core domain [1]. The β-sheet-rich insert is proposed to play a role in sequence-independent DNA and RNA binding [66]. The amino-terminal (D1) AAA+ modules of ClpB-type proteins are represented by a structure from T. thermophilus (right, PDB 1QVR). These proteins contain a long, leucine-rich coiled-coil propeller domain (pink) inserted into the small α-helical domain [67]. This propeller domain is proposed to play a role in interdomain communication and protein disaggregation, possibly acting as a molecular crowbar [67]. (b) Representative members of the HEC group [6]. The RFC1 AAA+ module from S. cerevisiae (center, PDB 1SXJ) represents a 'classical' AAA+ module containing no structural modifications and typifies the clamp loader clade to which it belongs [1,68]. The DnaA AAA+ module from Aquifex aeolicus (left, PDB 2HCB2HCB) contains an insert of two equal-sized helices (pink) after the second β-strand and is representative of the initiation clade [9]. The RuvB AAA+ module from T. thermophilus (right, PDB 1HQC) contains a β-hairpin insert (pink) between sensor 1 and its preceding helix [35]. This insert is characteristic of the RuvB family and is known to be important for the interaction of RuvB with RuvA in the resolution of Holliday junctions in DNA recombination [69,70]. The function of RuvB is discussed in the text. (c) Representatives of the PACTT group. Members of this group all contain a β-hairpin insert (cyan, shown in all three structures) between the sensor 1 strand and the preceding helix [1]. The BchI AAA+ module from Rhodobacter capsulatus Mg2+ chelatase (left, PDB 1G8P) belongs to the helix-2 insert clade. Members of this clade contain a small insert of two β-strands flanking a small α-helix (pink) in helix 2 of the αβα core domain and a long helical insert (yellow) between the fifth β-strand of the core domain and the small α-helical domain [1,24]. BchI proteins also contain a long, highly conserved β-hairpin insert (orange) upstream of the second β-strand of the core domain [24]. The function of BchI is discussed in the text. The carboxy-terminal ClpA AAA+ module (D2) from Escherichia coli (center, PDB 1KSF) [71] and the HslU AAA+ module from E. coli (right, PDB 1G4A) [72] are both representative members of the HCL clade, whose members are involved in protein unfolding and degradation. These structures contain an extended loop (pink) between the second core β-strand and the following helix [1] and a two or three stranded β-sheet insert (yellow) in the small α-helical domain of the AAA+ module, both characteristic of this clade. In addition, HslU family members contain an additional 130 amino acid I domain (orange, only part of the domain is resolved in the crystal structure) inserted into the core αβα domain of the AAA+ module, which is proposed to play a role in substrate recognition and unfolding [73].
Diversity of functional mechanisms
AAA+ proteins display a remarkable diversity of mechanisms of action. At the core of this diversity is the AAA+ molecular motor. ATP binding, hydrolysis and sensing are mediated by a number of different motifs and sequence elements within the AAA+ module, as outlined above (see Figure 1b). In general, hydrolysis is proposed to involve the abstraction of a proton from a molecule of water by the catalytic glutamate residue of the Walker B motif, thereby activating the water molecule for a subsequent nucleophilic attack on the γ-phosphate of bound ATP; the conserved lysine and serine/threonine residues of the Walker A motif act to bind the β- and γ-phosphates of the bound nucleotide and the Mg2+ ion, respectively [2, 4, 8, 9]. During the hydrolysis process, the amino-terminal and carboxy-terminal domains of the AAA+ module are proposed to move relative to one another, generating a mechanical force that can be used to affect remodeling events in associated molecules [4]. As most AAA+ proteins function as oligomeric assemblies, and as intersubunit communication exists between members of these assemblies, it is generally believed that nucleotide hydrolysis throughout the ring allows AAA+ assemblies to function in an efficient manner.
This central molecular motor has, however, been adapted to carry out an enormous variety of functions. This has generally been accomplished through direct structural modifications within the AAA+ module(s) themselves (Figure 2), and/or through the presence of additional domains at the amino and carboxyl termini of the AAA+ module(s) in a protein. In some cases, such additional domains are inserted within the AAA+ module. Hence, members of the AAA+ superfamily have evolved in such a way that they can recognize an enormous variety of different substrates and functional partners, thereby allowing the energy of nucleotide hydrolysis to be directed towards different remodeling events. Indeed, the various mechanisms employed by the AAA+ proteins are quite likely to be at least as diverse as the number of individual AAA+ families.
FtsH and ClpX
For example, both the FtsH and ClpX families of AAA+ proteins have similar roles in the cell in that they both function to unfold proteins and direct them for proteolytic degradation. Yet, the modifications and mechanisms by which they carry out these functions, while bearing some basic similarity, are quite distinct. Bacterial FtsH contains additional domains at the amino and carboxyl termini of its AAA+ module. The domain at the amino terminus acts to anchor the protein to the inner membrane of the bacterial cell, whereas the carboxy-terminal domain has protease activity [14, 15]. FtsH functions as a homohexamer and is proposed to consist of a combination of alternately 'open' (active) and 'closed' (inactive) subunits, which cycle in a nucleotide-dependent manner, driving the translocation of substrates to the protease active sites. It has been proposed that FtsH substrates pass via a tunnel along a 'closed' subunit into the protease active site of an adjacent 'open' subunit. Such a mechanism is consistent with the location of the proteolytic active sites at the periphery of the hexameric ring [15].
On the other hand, ClpX proteins form hexameric 'cap' structures that associate with a separate and structurally unrelated tetradecameric protease complex called ClpP [16, 17]. The ClpX hexamer uses the power of ATP hydrolysis to unfold protein substrates and direct them, through its central pore, into the proteolytic chamber of ClpP for degradation [18]. ClpP belongs to the family of self-compartmentalizing proteases that also includes the proteasome core particle. Interaction with the protease is via a conserved loop region within the AAA+ module of each subunit [19], whereas a zinc-binding domain at the amino terminus is important in binding protein factors that direct substrate specificity, substrate interaction and substrate translocation [20–22]. The zinc-binding domain of ClpX is proposed to undergo large nucleotide-dependent conformational changes in the course of its action [23].
Metal chelatases
Other AAA+ families have entirely different roles. The metal chelatase family, for example, is involved in the remodeling of small molecules. These enzymes utilize the power of ATP hydrolysis to insert Mg2+ or Co2+into porphyrin rings as part of the synthesis of (bacterio)chlorophyll or cobalamin (vitamin B12), respectively [24]. They work in conjunction with proteins containing von Willebrand factor type A (VWA) domains, which are metal-binding domains often involved in mediating protein-protein interactions [25]. Functional bacterial Mg2+ chelatase acts as a three-subunit enzyme consisting of an AAA+ module-containing subunit, BchI, and two other subunits, BchD (VWA containing) and BchH [26]. The BchD subunit contains an amino-terminal region similar to BchI and a carboxy-terminal VWA domain [24]. The amino-terminal region appears to represent a AAA+ module, but in many organisms key motifs are absent or disrupted, and the subunit has been shown to lack any independent ATPase activity [24, 27]. The BchH subunit has been shown to be responsible for porphyrin binding and to contain the chelatase active site, while the BchI and BchD subunits are proposed to power the chelation reaction and direct any necessary remodeling events [28].
Dynein
Yet another function of AAA+ proteins is exemplified by the dynein heavy chain (DHC) proteins of the PACCT group (Table 1). DHCs consist of a large amino-terminal stretch, six AAA+ modules fused together in tandem on a single polypeptide, an insertion between AAA+4 and AAA+5, and a carboxy-terminal domain [29]. DHCs are found throughout eukaryotes and serve as components of multimeric dynein complexes, where they function as molecular motors, using the hydrolysis of ATP as a source of energy for directing molecular motion and conformational changes. Different forms of DHCs and dynein complexes exist, including those associated with driving the motion of cilia and flagella, as well as cytoplasmic variants involved in the trafficking of various forms of 'cargo' molecules, such as vesicles, organelles and chromosomes along cytoskeletal filaments. Cytoplasmic dynein plays an important role in a variety of cellular processes including mitosis, nuclear envelope breakdown, retrograde vesicle transport, and maintenance of the Golgi apparatus, and has been shown to be essential for viability [29].
Electron microscopy studies have shown that DHC proteins fold into a globular 'head' structure, formed mainly by the six AAA+ modules, and two extensions, referred to as the stalk and the stem [30]. The stem is formed by the amino-terminal region of the protein and is responsible for protein-protein interaction/cargo binding [29]. The globular head is responsible for microtubule binding via a domain located at the tip of the structure [31]. The stalk and stem structures of the DHC molecules are highly flexible. Nucleotide binding, and possibly hydrolysis, bring about conformational changes in the globular head structure, as well as generating swing-like motions of the stem relative to the head and stalk. Although the mechanism of DHC function is not fully understood, it has been proposed that these swing-like motions of the stem effectively act as 'power-strokes', which allow translocation of dynein relative to microtubules associated with the stalk [30].
RuvB
The RuvB family of the HEC group (see Table 1) provides an example of AAA+ proteins acting on DNA. RuvB proteins are found throughout bacteria (see Table 1), and they play a key role in the later stages of homologous recombination. Homologous recombination is critically important in the maintenance of genome stability and repair of DNA damage, as well as in the generation of biological diversity. One important intermediate of this process is the Holliday junction, a DNA structure consisting of two homologous duplex DNA molecules associated via a single-stranded crossover. RuvB works together with the RuvA and RuvC proteins to process Holliday junctions into mature recombinant DNA molecules [32]. The RuvA protein forms tetrameric complexes which bind Holliday junctions with high affinity [33]. The RuvA protein interacts directly with the RuvB protein and facilitates its loading onto DNA [34].
RuvB proteins, which contain a single AAA+ module that is followed by a carboxy-terminal, winged-helix DNA-binding domain, have been shown by electron microscopy to bind to the Holliday junction as hexameric rings, contacting the bound RuvA on two opposite sides [35]. Together, the RuvA and RuvB proteins function as an ATP-dependent motor, promoting the branch migration of the Holliday junction, the process by which the junction moves along the DNA. RuvB molecules are proposed to act as ATP-driven pumps, driving helical rotation of double-stranded DNA, pulling it through the RuvA core and, thereby, promoting the branch migration process and increasing the formation of heteroduplex DNA. After branch migration, the junction is resolved by the action of the RuvC endonuclease, generating two recombinant DNA duplexes [32].
Other AAA+ families are involved in the remodeling of nucleic acids, or in the manipulation of entirely different classes of proteins. Clearly, the mechanisms employed by the AAA+ proteins are as diverse as the superfamily itself.
Despite the considerable increase in the number of proteins classified as belonging to the AAA+ superfamily and the extensive research that has been carried out on some of these proteins, much is yet to be learned about their functional mechanisms. The enormous size of the AAA+ superfamily, and the extensive functional diversity of its members, presents a challenging biological puzzle for those who wish to understand those molecular machines. As research continues, however, and additional insights are gained, we get closer to attaining a clearer picture of the remarkable process by which nature has adopted a single, common piece of molecular architecture for use in a vast array of cellular processes.
References
Iyer LM, Leipe DD, Koonin EV, Aravind L: Evolutionary history and higher order classification of AAA+ ATPases. J Struct Biol. 2004, 146: 11-31. 10.1016/j.jsb.2003.10.010.
Walker JE, Saraste M, Runswick MJ, Gay NJ: Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J. 1982, 1: 945-951.
Neuwald AF, Aravind L, Spouge JL, Koonin EV: A class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes. Genome Res. 1999, 9: 27-43.
Ogura T, Wilkinson AJ: AAA+ superfamily ATPases: common structure - diverse function. Genes Cells. 2001, 6: 575-597. 10.1046/j.1365-2443.2001.00447.x.
Erzberger JP, Berger JM: Evolutionary relationships and structural mechanisms of AAA+ proteins. Annu Rev Biophys Biomol Struct. 2006, 35: 93-114. 10.1146/annurev.biophys.35.040405.101933.
Ammelburg M, Frickey T, Lupas AN: Classification of AAA+ proteins. J Struct Biol. 2006, 156: 2-11.
Snider J, Houry WA: AAA+ proteins: diversity in function, similarity in structure. Biochem Soc Trans. 2008, 36: 72-77. 10.1042/BST0360072.
Leipe DD, Koonin EV, Aravind L: Evolution and classification of P-loop kinases and related proteins. J Mol Biol. 2003, 333: 781-815. 10.1016/j.jmb.2003.08.040.
Saraste M, Sibbald PR, Wittinghofer A: The P-loop - a common motif in ATP- and GTP-binding proteins. Trends Biochem Sci. 1990, 15: 430-434. 10.1016/0968-0004(90)90281-F.
Guenther B, Onrust R, Sali A, O'Donnell M, Kuriyan J: Crystal structure of the delta' subunit of the clamp-loader complex of E. coli DNA polymerase III. Cell. 1997, 91: 335-345. 10.1016/S0092-8674(00)80417-1.
Karata K, Inagawa T, Wilkinson AJ, Tatsuta T, Ogura T: Dissecting the role of a conserved motif (the second region of homology) in the AAA family of ATPases. Site-directed mutagenesis of the ATP-dependent protease FtsH. J Biol Chem. 1999, 274: 26225-26232. 10.1074/jbc.274.37.26225.
Ogura T, Whiteheart SW, Wilkinson AJ: Conserved arginine residues implicated in ATP hydrolysis, nucleotide-sensing, and inter-subunit interactions in AAA and AAA+ ATPases. J Struct Biol. 2004, 146: 106-112. 10.1016/j.jsb.2003.11.008.
Johnson A, O'Donnell M: Ordered ATP hydrolysis in the gamma complex clamp loader AAA+ machine. J Biol Chem. 2003, 278: 14406-14413. 10.1074/jbc.M212708200.
Tomoyasu T, Yamanaka K, Murata K, Suzaki T, Bouloc P, Kato A, Niki H, Hiraga S, Ogura T: Topology and subcellular localization of FtsH protein in Escherichia coli. J Bacteriol. 1993, 175: 1352-1357.
Suno R, Niwa H, Tsuchiya D, Zhang X, Yoshida M, Morikawa K: Structure of the whole cytosolic region of ATP-dependent protease FtsH. Mol Cell. 2006, 22: 575-585. 10.1016/j.molcel.2006.04.020.
Grimaud R, Kessel M, Beuron F, Steven AC, Maurizi MR: Enzymatic and structural similarities between the Escherichia coli ATP-dependent proteases, ClpXP and ClpAP. J Biol Chem. 1998, 273: 12476-12481. 10.1074/jbc.273.20.12476.
Ortega J, Singh SK, Ishikawa T, Maurizi MR, Steven AC: Visualization of substrate binding and translocation by the ATP-dependent protease, ClpXP. Mol Cell. 2000, 6: 1515-1521. 10.1016/S1097-2765(00)00148-9.
Zolkiewski M: A camel passes through the eye of a needle: protein unfolding activity of Clp ATPases. Mol Microbiol. 2006, 61: 1094-1100. 10.1111/j.1365-2958.2006.05309.x.
Kim YI, Levchenko I, Fraczkowska K, Woodruff RV, Sauer RT, Baker TA: Molecular determinants of complex formation between Clp/Hsp100 ATPases and the ClpP peptidase. Nat Struct Biol. 2001, 8: 230-233. 10.1038/84967.
Singh SK, Rozycki J, Ortega J, Ishikawa T, Lo J, Steven AC, Maurizi MR: Functional domains of the ClpA and ClpX molecular chaperones identified by limited proteolysis and deletion analysis. J Biol Chem. 2001, 276: 29420-29429. 10.1074/jbc.M103489200.
Wojtyra UA, Thibault G, Tuite A, Houry WA: The N-terminal zinc binding domain of ClpX is a dimerization domain that modulates the chaperone function. J Biol Chem. 2003, 278: 48981-48990. 10.1074/jbc.M307825200.
Thibault G, Yudin J, Wong P, Tsitrin V, Sprangers R, Zhao R, Houry WA: Specificity in substrate and cofactor recognition by the N-terminal domain of the chaperone ClpX. Proc Natl Acad Sci USA. 2006, 103: 17724-17729. 10.1073/pnas.0601505103.
Thibault G, Tsitrin Y, Davidson T, Gribun A, Houry WA: Large nucleotide-dependent movement of the N-terminal domain of the ClpX chaperone. EMBO J. 2006, 25: 3367-3376. 10.1038/sj.emboj.7601223.
Fodje MN, Hansson A, Hansson M, Olsen JG, Gough S, Willows RD, Al-Karadaghi S: Interplay between an AAA module and an integrin I domain may regulate the function of magnesium chelatase. J Mol Biol. 2001, 311: 111-122. 10.1006/jmbi.2001.4834.
Whittaker CA, Hynes RO: Distribution and evolution of von Wille-brand/integrin A domains: widely dispersed domains with roles in cell adhesion and elsewhere. Mol Biol Cell. 2002, 13: 3369-3387. 10.1091/mbc.E02-05-0259.
Walker CJ, Willows RD: Mechanism and regulation of Mg-chelatase. Biochem J. 1997, 327: 321-333.
Sirijovski N, Olsson U, Lundqvist J, Al-Karadaghi S, Willows RD, Hansson M: ATPase activity associated with the magnesium chelatase H-subunit of the chlorophyll biosynthetic pathway is an artefact. Biochem J. 2006, 400: 477-484. 10.1042/BJ20061103.
Reid JD, Hunter CN: Current understanding of the function of magnesium chelatase. Biochem Soc Trans. 2002, 30: 643-645. 10.1042/BST0300643.
Sakato M, King SM: Design and regulation of the AAA+ microtubule motor dynein. J Struct Biol. 2004, 146: 58-71. 10.1016/j.jsb.2003.09.026.
Burgess SA, Walker ML, Sakakibara H, Oiwa K, Knight PJ: The structure of dyneinc by negative stain electron microscopy. J Struct Biol. 2004, 146: 205-216. 10.1016/j.jsb.2003.10.005.
Gee MA, Heuser JE, Vallee RB: An extended microtubule-binding structure within the dynein motor domain. Nature. 1997, 390: 636-639. 10.1038/37663.
West SC: Processing of recombination intermediates by the RuvABC proteins. Annu Rev Genet. 1997, 31: 213-244. 10.1146/annurev.genet.31.1.213.
Iwasaki H, Takahagi M, Nakata A, Shinagawa H: RuvA and RuvB proteins specifically interact with Holliday junctions and promote branch migration. Genes Dev. 1992, 6: 2214-2220. 10.1101/gad.6.11.2214.
Parsons CA, West SC: Formation of a RuvAB-Holliday junction complex in vitro. J Mol Biol. 1993, 232: 397-405. 10.1006/jmbi.1993.1399.
Yamada K, Kunishima N, Mayanagi K, Ohnishi T, Nishino T, Iwasaki H, Shinagawa H, Morikawa K: Crystal structure of the Holliday junction migration motor protein RuvB from Thermus thermophilus HB8. Proc Natl Acad Sci USA. 2001, 98: 1442-1447. 10.1073/pnas.031470598.
Ito K, Akiyama Y: Cellular functions, mechanism of action, and regulation of FtsH protease. Annu Rev Microbiol. 2005, 59: 211-231. 10.1146/annurev.micro.59.030804.121316.
Baas PW, Karabay A, Qiang L: Microtubules cut and run. Trends Cell Biol. 2005, 15: 518-524. 10.1016/j.tcb.2005.08.004.
Whiteheart SW, Matveeva EA: Multiple binding proteins suggest diverse functions for the N-ethylmaleimide sensitive factor. J Struct Biol. 2004, 146: 32-43. 10.1016/j.jsb.2003.09.015.
Jentsch S, Rumpf S: Cdc48 (p97): a "molecular gearbox" in the ubiquitin pathway?. Trends Biochem Sci. 2007, 32: 6-11. 10.1016/j.tibs.2006.11.005.
Thoms S, Erdmann R: Peroxisomal matrix protein receptor ubiquitination and recycling. Biochim Biophys Acta. 2006, 1763: 1620-1628. 10.1016/j.bbamcr.2006.08.046.
Cruciat CM, Hell K, Folsch H, Neupert W, Stuart RA: Bcs1p, an AAA-family member, is a chaperone for the assembly of the cytochrome bc(1) complex. EMBO J. 1999, 18: 5226-5233. 10.1093/emboj/18.19.5226.
Smith DM, Benaroudj N, Goldberg A: Proteasomes and their associated ATPases: a destructive combination. J Struct Biol. 2006, 156: 72-83.
Portis AR: Rubisco activase - Rubisco's catalytic chaperone. Photosynth Res. 2003, 75: 11-27. 10.1023/A:1022458108678.
Gribun A, Cheung KL, Huen J, Ortega J, Houry WA: Yeast Rvb1 and Rvb2 are ATP-dependent DNA helicases that form a heterohexameric complex. J Mol Biol. 2008, 376: 1320-1333. 10.1016/j.jmb.2007.12.049.
Fan N, Cutting S, Losick R: Characterization of the Bacillus subtilis sporulation gene spoVK. J Bacteriol. 1992, 174: 1053-1054.
Drescher A, Ruf S, Calsa T, Carrer H, Bock R: The two largest chloroplast genome-encoded open reading frames of higher plants are essential genes. Plant J. 2000, 22: 97-104. 10.1046/j.1365-313x.2000.00722.x.
Lee YJ, Wickner RB: AFG1, a new member of the SEC18-NSF, PAS1, CDC48-VCP, TBP family of ATPases. Yeast. 1992, 8: 787-790. 10.1002/yea.320080912.
Johnson A, O'Donnell M: Cellular DNA replicases: components and dynamics at the replication fork. Annu Rev Biochem. 2005, 74: 283-315. 10.1146/annurev.biochem.73.011303.073859.
Hishida T, Ohya T, Kubota Y, Kamada Y, Shinagawa H: Functional and physical interaction of yeast Mgs1 with PCNA: impact on RAD6-dependent DNA damage tolerance. Mol Cell Biol. 2006, 26: 5509-5517. 10.1128/MCB.00307-06.
Schaeffer PM, Headlam MJ, Dixon NE: Protein-protein interactions in the eubacterial replisome. IUBMB Life. 2005, 57: 5-12. 10.1080/15216540500058956.
Davey MJ, Jeruzalmi D, Kuriyan J, O'Donnell M: Motors and switches: AAA+ machines within the replisome. Nat Rev Mol Cell Biol. 2002, 3: 826-835. 10.1038/nrm949.
West SC: Processing of recombination intermediates by the RuvABC proteins. Annu Rev Genet. 1997, 31: 213-244. 10.1146/annurev.genet.31.1.213.
Schmid S, Seitz T, Haas D: Cointegrase, a naturally occurring, truncated form of IS21 transposase, catalyzes replicon fusion rather than simple insertion of IS21. J Mol Biol. 1998, 282: 571-583. 10.1006/jmbi.1998.2041.
Pham P, Frei KP, Woo W, Truong DD: Molecular defects of the dystonia-causing torsinA mutation. Neuroreport. 2006, 17: 1725-1728. 10.1097/WNR.0b013e3280101220.
Rotanova TV, Botos I, Melnikov EE, Rasulova F, Gustchina A, Maurizi MR, Wlodawer A: Slicing a protease: structural features of the ATP-dependent Lon proteases gleaned from investigations of isolated domains. Protein Sci. 2006, 15: 1815-1828. 10.1110/ps.052069306.
Maiorano D, Lutzmann M, Mechali M: MCM proteins and DNA replication. Curr Opin Cell Biol. 2006, 18: 130-136. 10.1016/j.ceb.2006.02.006.
Bourniquel AA, Bickle TA: Complex restriction enzymes: NTP-driven molecular motors. Biochimie. 2002, 84: 1047-1059. 10.1016/S0300-9084(02)00020-2.
Galani K, Nissan TA, Petfalski E, Tollervey D, Hurt E: Rea1, a dynein-related nuclear AAA-ATPase, is involved in late rRNA processing and nuclear export of 60 S subunits. J Biol Chem. 2004, 279: 55411-55418. 10.1074/jbc.M406876200.
Snider J, Houry WA: MoxR AAA+ ATPases: a novel family of molecular chaperones?. J Struct Biol. 2006, 156: 200-209.
Schumacher J, Joly N, Rappas M, Zhang X, Buck M: Structures and organisation of AAA+ enhancer binding proteins in transcriptional activation. J Struct Biol. 2006, 156: 190-199.
Gwinn ML, Ramanathan R, Smith HO, Tomb JF: A new transformation-deficient mutant of Haemophilus influenzae Rd with normal DNA uptake. J Bacteriol. 1998, 180: 746-748.
Schoenhofen IC, Li G, Strozen TG, Howard SP: Purification and characterization of the N-terminal domain of ExeA: a novel ATPase involved in the type II secretion pathway of Aeromonas hydrophila. J Bacteriol. 2005, 187: 6370-6378. 10.1128/JB.187.18.6370-6378.2005.
Leipe DD, Koonin EV, Aravind L: STAND, a class of P-loop NTPases including animal and plant regulators of programmed cell death: multiple, complex domain architectures, unusual phyletic patterns, and evolution by horizontal gene transfer. J Mol Biol. 2004, 343: 1-28. 10.1016/j.jmb.2004.08.023.
Sousa MC, Kessler BM, Overkleeft HS, McKay DB: Crystal structure of HslUV complexed with a vinyl sulfone inhibitor: corroboration of a proposed mechanism of allosteric activation of HslV by HslU. J Mol Biol. 2002, 318: 779-785. 10.1016/S0022-2836(02)00145-6.
PyMOL. [http://pymol.sourceforge.net]
Matias PM, Gorynia S, Donner P, Carrondo MA: Crystal structure of the human AAA+ protein RuvBL1. J Biol Chem. 2006, 281: 38918-38929. 10.1074/jbc.M605625200.
Lee S, Sowa ME, Watanabe YH, Sigler PB, Chiu W, Yoshida M, Tsai FT: The structure of ClpB: a molecular chaperone that rescues proteins from an aggregated state. Cell. 2003, 115: 229-240. 10.1016/S0092-8674(03)00807-9.
Bowman GD, O'Donnell M, Kuriyan J: Structural analysis of a eukaryotic sliding DNA clamp-clamp loader complex. Nature. 2004, 429: 724-730. 10.1038/nature02585.
Han YW, Iwasaki H, Miyata T, Mayanagi K, Yamada K, Morikawa K, Shinagawa H: A unique beta-hairpin protruding from AAA+ ATPase domain of RuvB motor protein is involved in the interaction with RuvA DNA recognition protein for branch migration of Holliday junctions. J Biol Chem. 2001, 276: 35024-35028. 10.1074/jbc.M103611200.
Yamada K, Miyata T, Tsuchiya D, Oyama T, Fujiwara Y, Ohnishi T, Iwasaki H, Shinagawa H, Ariyoshi M, Mayanagi K, Morikawa K: Crystal structure of the RuvA-RuvB complex: a structural basis for the Holliday junction migrating motor machinery. Mol Cell. 2002, 10: 671-681. 10.1016/S1097-2765(02)00641-X.
Guo F, Maurizi MR, Esser L, Xia D: Crystal structure of ClpA, an Hsp100 chaperone and regulator of ClpAP protease. J Biol Chem. 2002, 277: 46743-46752. 10.1074/jbc.M207796200.
Wang J, Song JJ, Franklin MC, Kamtekar S, Im YJ, Rho SH, Seong IS, Lee CS, Chung CH, Eom SH: Crystal structures of the HslVU peptidase-ATPase complex reveal an ATP-dependent proteolysis mechanism. Structure. 2001, 9: 177-184. 10.1016/S0969-2126(01)00570-6.
Groll M, Bochtler M, Brandstetter H, Clausen T, Huber R: Molecular machines for protein degradation. ChemBioChem. 2005, 6: 222-256. 10.1002/cbic.200400313.
Acknowledgements
JS and GT held graduate fellowships from the Natural Sciences and Engineering Research Council of Canada. GT also received a graduate fellowship from the Fonds Quebecois de la Recherche sur la Nature et les Technologies. This work has been supported in part by the Natural Sciences and Engineering Research Council of Canada Discovery Grant Program to WAH.
Author information
Authors and Affiliations
Corresponding author
Additional information
Jamie Snider, Guillaume Thibault contributed equally to this work.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
About this article
Cite this article
Snider, J., Thibault, G. & Houry, W.A. The AAA+ superfamily of functionally diverse proteins. Genome Biol 9, 216 (2008). https://doi.org/10.1186/gb-2008-9-4-216
Published:
DOI: https://doi.org/10.1186/gb-2008-9-4-216